Kinase Selectivity Profiling That Shows You More Than % Inhibition
KinSight™ profiles your compounds against 400+ validated kinases using continuous, real-time assays — delivering true selectivity data, TDI detection, and MOA insights that endpoint platforms systematically miss.
400+
Wild-type protein kinases
2
Week report turnaround
n=2
All assays run in duplicate
NCI Funded
MIT-licensed technology
Why Settle for a Single Snapshot of Enzyme Activity?
Most kinome profiling platforms take one measurement and call it done. But a single time point assumes the reaction is linear, stable, and uninhibited at that exact moment — an assumption that silently breaks down more often than you'd think.
The example below is real: four kinases that look nearly identical at endpoint. Progress curves tell a completely different story.
Linear Kinetics
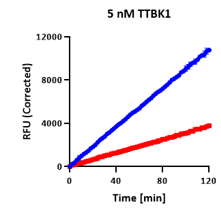
Endpoint and kinetics fully agree.
Two clean straight lines from origin — DMSO rising steeply, compound shallow but linear. This is a straightforward reversible inhibitor, and the only pattern endpoint profiling reliably identifies correctly.
Initial Lag
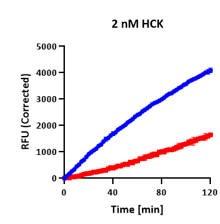
Both curves flat ~20 min before rate establishes.
The enzyme takes time to reach its steady-state activity. Endpoint assays that sample during this lag calculate a falsely low rate, inflating apparent inhibition. Continuous assays identify the true linear window and calculate from there.
TDI Detected
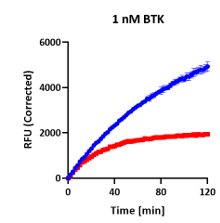
Inhibition deepens over time — slope progressively decelerates.
Both curves start from a shared elevated baseline, then diverge as the compound progressively occupies or modifies the enzyme. A single endpoint measurement captures only a snapshot — and misses the mechanism and its true potency entirely.
Assay Artifact
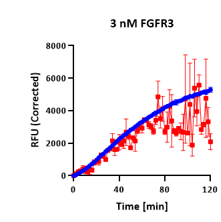
Heavy scatter throughout — noise invisible at endpoint.
The compound curve shows extreme replicate variability. The 61% inhibition read is an artifact with no statistical basis. Endpoint platforms report this with the same confidence as a clean result.

Endpoint and kinetics fully agree.
Two clean straight lines from origin — DMSO rising steeply, compound shallow but linear. This is a straightforward reversible inhibitor, and the only pattern endpoint profiling reliably identifies correctly.

Both curves flat ~20 min before rate establishes.
The enzyme takes time to reach its steady-state activity. Endpoint assays that sample during this lag calculate a falsely low rate, inflating apparent inhibition. Continuous assays identify the true linear window and calculate from there.

Inhibition deepens over time — slope progressively decelerates.
Both curves start from a shared elevated baseline, then diverge as the compound progressively occupies or modifies the enzyme. A single endpoint measurement captures only a snapshot — and misses the mechanism and its true potency entirely.

Heavy scatter throughout — noise invisible at endpoint.
The compound curve shows extreme replicate variability. The 61% inhibition read is an artifact with no statistical basis. Endpoint platforms report this with the same confidence as a clean result.
Continuous profiling does more than rank kinases by % inhibition — it reveals whether the assay is linear, noisy, delayed, or time-dependent. That's the difference between data you can act on and data that looks right until it isn't.
Four Steps From Compound to Kinetic Insights
A streamlined process designed to get you high-confidence profiling data with minimal friction — from submission to annotated report in two weeks.
1. Submit your compound and select a panel
Choose from six predefined panels or work with our team to build a custom target list. Single concentration or dose-response available.
2. We run continuous assays at your ATP conditions
ATP Km for maximal potency insight, 1 mM for physiological relevance, or both. Every well generates a full progress curve — actual rates from multiple data points.
3. Expert review and annotation
Our scientists review every result and annotate TDI signals, unusual kinetics, and MOA insights. Not just a data dump — a report you can act on.
4. Full data package delivered in two weeks
Waterfall plots, kinome tree visualization, tabular % inhibition data, and annotated progress curves — ready to share with your team and advance your program.
Ready to Profile Your Compound?
Tell us about your compound and goals. We'll recommend the right panel and ATP conditions — or scope the full project with one of our scientists directly.
Most projects start with a 20-minute scoping call. No commitment required.
Kinase Selectivity Panels
Panel Name |
Panel Size |
ATP Options |
Target List |
|
Wild Type Protein Kinase Panel |
401 |
ATP Km and 1mM
now offering custom ATP concentrations*
|
|
|---|---|---|---|
|
Full Protein Kinase Panel |
450 |
ATP Km and 1mM
now offering custom ATP concentrations*
|
View Panel > |
|
Tyrosine Kinase (TK) Panel |
83 |
ATP Km and 1mM
now offering custom ATP concentrations*
|
View Panel > |
|
Cyclin Dependent Kinase (CDK) Panel |
32 |
ATP Km and 1mM
now offering custom ATP concentrations*
|
View Panel > |
|
Mutant Protein Kinase Panel |
53 |
ATP Km and 1mM
now offering custom ATP concentrations*
|
View Panel > |
|
KiNode Panel |
69 |
ATP Km and 1mM
now offering custom ATP concentrations*
|
View Panel > |
|
Custom Panel |
1-100 |
ATP Km and 1mM
now offering custom ATP concentrations*
|
Contact Us to Design > |
*Kinase selectivity profiling is conducted on a bi-monthly schedule, with assays run at the apparent ATP Km (to enable accurate potency assessment) or at 1 mM ATP (to approximate physiological conditions). Alternative ATP concentrations can be accommodated upon request.
Why Researchers Choose PhosphoSens® Over Endpoint Methods
Most kinase assays give you a single data point at a single moment in time. PhosphoSens generates a continuous activity trace — so you see the full kinetics, not just a snapshot.
True initial rates, no artifacts
Calculate rates from the true linear region of the progress curve — free from lag phase, substrate depletion, or enzyme instability artifacts.
Detect TDI and slow-binding inhibitors
Capture time-dependent mechanisms — TDI, slow-on/slow-off, irreversible binding — that endpoint assays miss entirely.
Physiologically relevant conditions
Compatible with physiological ATP, Mg²⁺, Mn²⁺, and Ca²⁺. Validated substrates with defined phospho-acceptor sites — no antibody cross-reactivity.
| Platform | Measures activity directly | Detects time dependent inhibition | Real-time kinetic readout | Detects all inhibitor types |
|---|---|---|---|---|
| AssayQuant PhosphoSens® | ||||
| Luminescent assayse.g. Kinase-Glo, ADP-Glo | ||||
| Radiometric assayse.g. ³³P/³²P filter binding | ||||
| Mobility shift assayse.g. Caliper EZ Reader | ||||
| Competition bindinge.g. KINOMEscan, KdELECT |
✓ Supported | Indirect = proxy readout, endpoint assumptions apply | ✗ Not supported
Read: Continuous vs. Endpoint Kinase Assays →Kinase Activity Directly From Cell Lysates
Skip the purified enzyme. PhosphoSens-Lysate assays measure kinase activity in native cellular context — capturing the biology your recombinant assays can't.
Continuous Phosphatase Activity Assays
The same real-time, direct measurement you rely on for kinases — now for phosphatases. 50+ validated targets, same PhosphoSens® platform.
Resources for Kinase Assay Scientists
Deep Kinetic Characterization: When IC50 Isn't Enough
A continuous assay, like our PhosphoSens assay, that measures the catalytic event of phosphorylation in real time enables determination of kinetic parameters that power your drug discovery research.
Continuous vs. Endpoint Kinase Assays: What You Need to Know
PhosphoSens-Kinetic Phosphatase Inhibitor IC50 Determination
Frequently Asked Questions
Your pain? We understand. This is why we do what we do, and can provide you with an experience like no other.
What is a PhosphoSens® kinase assay?
PhosphoSens® assays are continuous, real-time kinase activity assays that directly measure phosphorylation of a substrate peptide throughout the reaction. Unlike endpoint assays that capture a single time point, PhosphoSens generates a full progress curve — enabling true kinetic analysis including IC₅₀, Kᵢ, kobs, and time-dependent inhibition (TDI) from a single experiment.
What's the difference between PhosphoSens-Kinetic and PhosphoSens-Red?
PhosphoSens-Kinetic is a continuous fluorescence assay that monitors kinase activity in real time throughout the reaction, generating full progress curves. PhosphoSens-Red is a time-resolved fluorescence (TRF) endpoint format optimized for higher throughput screening. Both use the same underlying PhosphoSens® substrate technology — the choice depends on whether you need kinetic depth (Kinetic) or screening throughput (Red).
How does a continuous kinase assay detect time-dependent inhibition (TDI)?
TDI compounds produce a characteristic change in the progress curve shape — the inhibition deepens over time as the compound slowly occupies or covalently modifies the enzyme. Because PhosphoSens monitors activity continuously, this curve deviation is directly visible. Endpoint assays that measure at a single time point will either miss TDI entirely or mischaracterize its potency, depending on when they sample the reaction.
What plate reader is required for PhosphoSens assays?
PhosphoSens-Kinetic assays require a standard fluorescence plate reader capable of kinetic reads (repeated measurements over time) with excitation/emission appropriate for the Sox fluorophore. PhosphoSens-Red assays require a time-resolved fluorescence (TRF) reader. Most modern multimode readers in drug discovery labs are compatible. Contact us if you need compatibility confirmation for your specific instrument.
Can't find my target in the catalog?
We offer custom assay development for kinase targets not currently in our catalog. Our team can design and validate a PhosphoSens substrate for your target, typically within 8–12 weeks. Contact us to discuss your target, timeline, and project requirements.
Ready to Characterize Your Compound?
Talk to one of our scientists about your target, your workflow, and which assay format will get you the data you need.
Stay Informed
Want to hear the latest about our technology? Be among the first to learn about our latest products and services.